RG Ziemert
Translational Genome Mining for Natural Products
The Ziemert lab is interested in the evolution and distribution of bacterial secondary metabolites. These bioactive compounds are especially important in human medicine as the chemical scaffolds are the main templates for new drugs such as antibiotics and anticancer agents. We are a highly interdisciplinary and enthusiastic group at the interface of bioinformatics, molecular and environmental microbiology, and phylogenetics. We are currently developing bioinformatic tools for the discovery of natural products from microbial genomes, elucidating the evolutionary history of their biosynthetic gene clusters and tracing their distribution in the environment.
Microbial Natural Products
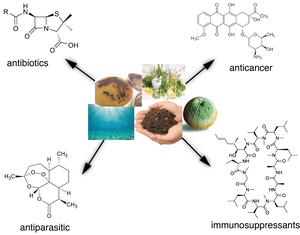
Natural products are small bioactive molecules produced by living organisms that have a tremendous impact on drug discovery and human health. Half of all anticancer drugs and 66% of all antibacterial drugs approved over the last 30 years are natural products or natural product derivatives. Of the 12 antibacterial classes, nine are derived from a natural product template. Widely-prescribed antibacterial and chemotherapeutic drugs like erythromycin, streptomycin, penicillin, tetracycline, vancomycin, and doxorubicin all come from Nature and a large part of them from microorganisms. More precisely, 90% of all antibiotics in clinical use are derived from microorganisms. With the ever-increasing rise of multi-resistant bacteria and reemerging infectious diseases we are facing a global health crisis and find ourselves in urgent need for new classes of therapeutic drugs including new antibiotics.
Genome Mining
The revolution of next generation sequencing methods and along with it the enormous amount of publicly available DNA sequences, has now shifted traditional bioactivity guided or chemical screening methods towards genomic based approaches and resulted in a renaissance of natural product research. Mining microbial genomes for their biosynthetic gene clusters revealed the true potential of many organisms to produce natural products. Even well probed genera such as Streptomyces were shown to be able to produce on average 10 times more chemical compounds than have been isolated under laboratory conditions. Moreover, massive whole genome sequencing projects demonstrated that many more species and environments are encoding novel chemical diversity and more compounds are waiting for discovery. Now the challenge is to effectively mine the huge amount of available data, prioritize the most promising gene clusters and develop effective ways to detect and elucidate encoded compounds.
Natural Products in the Environment
Natural products are often produced as secondary metabolites by plants, fungi and bacteria. Secondary metabolites are specialized compounds that are distinguished from primary metabolites because they are not directly involved in cell growth or reproduction, are usually taxonomically restricted, and have specialized but nonetheless important functions including signaling, nutrient acquisition and defense. For the vast majority of bacterial natural products known so far it is not known why and when they are produced in the environment. However, the examples that are known, prove that natural products are important compounds that bacteria use to sense and respond to their environment and that their exact ecological functions are as diverse as their chemical structures.
Evolution of biosynthetic gene clusters and their producers
Natural products are results of millions of years of evolution that shaped and refined biosynthetic processes. Studying the evolutionary processes that shaped biosynthetic gene clusters as they can be seen today, we are able to support drug discovery on various levels:
I) identification of promising producer organisms, II) prediction of molecular (sub-) structures based on their genetic code, and III) synthetic biology for designing new and optimizing known compounds.
Our Focus
The Ziemert lab is using various genome mining techniques to detect and identify natural product gene clusters in bacteria and environments. Hereby we are not only interested in finding new bioactive molecules for drug discovery, but also aim to elucidate when and by whom are these molecules produced, how are they vertically and horizontally transferred and how exactly is this huge chemical diversity evolving in nature. We are focusing on developing more effective and targeted genome mining approaches and develop bioinformatic webtools for the natural product community.
