ARTS – Antibiotic Resistant Target Seeker
Click here for the ARTS Homepage
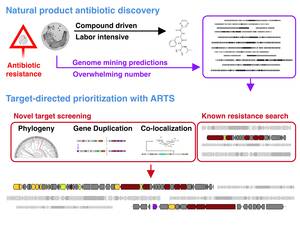
The ARTS webserver provides an orthogonal approach to uncovering antibiotic potential in genomic data by highlighting known and putative resistance genes. In combination with antiSMASH, ARTS serves to prioritize biosynthetic gene cluster (BGC) for those that contain resistance genes. In the light of many examples of co-localization of resistance genes in antibiotic gene clusters, this results in guided wet-lab experiments to discover those likely to produce antibiotics. Additionally, the gene annotation and target prediction can give insight into the mode of action to further accelerate investigation.
Reference
The Antibiotic Resistant Target Seeker (ARTS), an exploration engine for antibiotic cluster prioritization and novel drug target discovery.
Alanjary M, Kronmiller B, Adamek M, Blin K, Weber T, Huson D, Philmus B, Ziemert N. Nucleic Acids Res. 2017 May 2. doi: 10.1093/nar/gkx360
