NaPDoS - Natural Product Domain seeker
Click here for the NaPDoS Homepage
NaPDos is a bioinformatic tool for the rapid detection and analysis of secondary metabolite genes. This tool is designed to detect and extract C- and KS- domains from DNA or amino acid sequence data, including PCR amplicon products, individual genes, whole genomes, and metagenomic data sets.
Candidate secondary metabolite domains are identified by sequence comparison to a broad set of manually curated reference genes from well-characterized chemical pathways. Candidate gene sequences are extracted, trimmed, translated (if necessary) and subjected to domain-specific phylogenetic clustering to predict what their putative products might be, and to determine whether these products are likely to produce compounds similar to or different from previously known biosynthetic pathways. Because NaPDoS analyzes small but phylogenetically conserved domains of usually very large secondary metabolite gene clusters, it is specifically useful for metagenomic data and PCR amplicons.
Reference
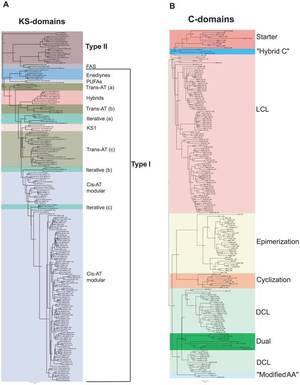
The natural product domain seeker NaPDoS: a phylogeny based bioinformatic tool to classify secondary metabolite gene diversity. Ziemert N, Podell S, Penn K, Badger JH, Allen E, Jensen PR. PLoS One. 2012;7(3):e34064. doi: 10.1371/journal.pone.0034064. Epub 2012 Mar 29.
