News
16.05.2023
Reconstruction, Comparison and Growth Simulation of Strain-specific Metabolic Computer Models of Staphylococcus haemolyticus
Gwendolyn O. Gusak is developing a collection of high-quality models for S. haemolyticus in her master’s thesis.
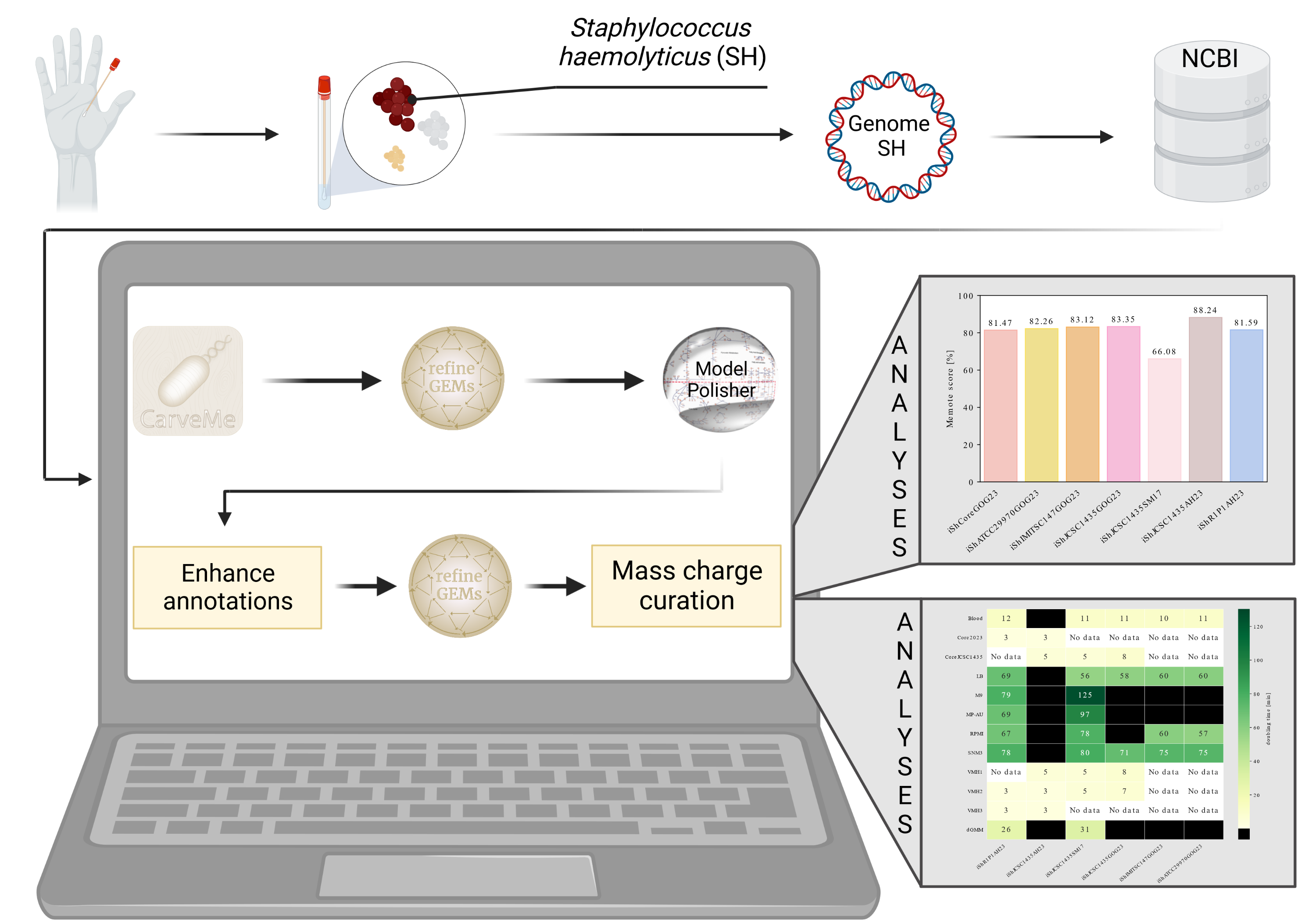
In times of increasingly spreading antibiotic resistance, the comprehensive investigation of opportunistic pathogens is becoming more and more critical. So-called in-sillico approaches (i.e., biological experiments carried out on the computer), such as the simulation of strain-specific metabolic models on the genome level (GEMs), make an essential contribution to investigating pathogens, as they enable the rapid testing of plausible laboratory scenarios. As part of her master’s thesis, “Reconstruction of strain-specific metabolic models of Staphylococcus haemolyticus,” Ms. Gwendolyn O. Gusak developed three new GEMs of the organism Staphylococcus haemolyticus (SH) and compared them to GEMs automatically generated for the same organism. Their new workflow from genome to draft GEM shows good results, as 98.69% of all protein sequences obtained valid identifiers from the NCBI database. Her resulting model iShIMITSC147GOG23 showed similar results compared to SH’s other models. In addition, as part of her work, she extended the script collection called refineGEMs to include an algorithm that fills gaps in the models, which she plans to continue working on after completing her thesis. Comparing the MEMOTE scores of the new models with the automatically generated models shows that the quality of the models is the same, except for the oldest model from 2017, which has a low MEMOTE score of 66.08%. The results of their growth simulations mostly show doubling times in a plausible range for laboratory research. She makes her work on the software and the models themselves publicly available on GitHub and in the BioModels database common to systems biology. After completing this work, Ms. Gusak plans to have her results peer-reviewed and published by a journal.